Review Article
MicroRNAs as Master Regulators of Plant Development and Stress Adaptation
Vasudeva Reddy Netala1, Nora M Al Aboud2 and Vijaya Tartte3
1School of Chemical Engineering and Technology, North University of China, Taiyuan, 030051, China.
2Department of Biology, Faculty of Science, Umm Al-Qura University, Makkah, Saudi Arabia
3Department of Biotechnology, Sri Venkateswara University, Tirupati, Andhra Pradesh, 517502, India.
2Department of Biology, Faculty of Science, Umm Al-Qura University, Makkah, Saudi Arabia
3Department of Biotechnology, Sri Venkateswara University, Tirupati, Andhra Pradesh, 517502, India.
*Corresponding author:Vasudeva Reddy Netala, Department of Chemical Engineering and Technology, North University of China, Taiyuan, 030051, China. E-mail Id: drreddy0205@qq.com
Copyright: © Netala VR, et al. 2025. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
Article Information:Submission: 20/05/2025; Accepted: 18/06/2025; Published: 21/06/2025
Abstract
MicroRNAs (miRNAs) are small (~20–24 nt), non-coding RNAs that serve as master regulators of gene expression in plants, playing pivotal roles in growth, development, and stress adaptation. This review systematically examines the molecular mechanisms of miRNA biogenesis, from
transcription and processing to their integration into RNA-induced silencing complexes (RISCs). We highlight their spatiotemporal regulation of key developmental processes—including leaf morphogenesis, root architecture, phase transitions, and reproductive development—and their adaptive roles in abiotic (drought, salinity, nutrient deficiency) and biotic (pathogens, herbivores) stress responses. The evolutionary conservation of miRNA pathways across plant species underscores their functional significance, while emerging biotechnological applications, such as engineered miRNAs and CRISPR-based editing, offer innovative strategies for crop improvement. By synthesizing current advances and future
perspectives (e.g., single-cell miRNAomics, synthetic networks, and cross-kingdom signaling), this review provides a comprehensive framework for understanding miRNA-mediated regulation in plants and its potential to address global agricultural challenges.
Keywords:MicroRNAs (miRNAs); miRNA biogenesis; Gene regulation; Plant development; Stress responses;Crop improvement;Evolutionary conservation; RNA interference (RNAi)
Abbreviations
miRNAs: MicroRNAs; RNAi: RNA interference; pri-miRNAs:
Primary miRNAs; DCL: DICER-LIKE Protein; RISC: RNAinduced
silencing complex; AGO: Argonaute; TFs: Transcription
Factors; ceRNAs: Competing endogenous RNAs; HATs: Histone
acetyltransferases; HMTs: Histone methyltransferases; PRC2:
Polycomb Repressive Complex 2; ADARs: Adenosine deaminases;
PTI: PAMP-triggered immunity; SA: Salicylic acid: JA: Jasmonic acid;
RBPs: RNA-Binding Proteins; ARFs: AUXIN RESPONSE FACTORs;
SPL: SQUAMOSA PROMOTER.
BINDING PROTEIN-LIKE:
AP2: APETALA2; SAM:Shoot Apical Meristem; AM:
Axillary Meristem; LCR: LEAF CURLING RESPONSIVENESS;
WUS:WUSCHEL; CLV3:CLAVATA3; PR: Pathogenesis-Related.Introduction
MicroRNAs (miRNAs) are small, non-coding RNA molecules
that typically range from 20 to 24 nucleotides in length, and they play
a pivotal role in the post-transcriptional regulation of gene expression
in plants [1]. These highly conserved regulatory molecules function
as critical modulators of cellular processes, exerting their influence
through sequence-specific interactions with target messenger RNAs
[2]. Since their initial discovery in the early 1990s, miRNAs have
been recognized as master regulators that orchestrate a wide array of
biological functions throughout a plant’s life cycle. Their importance
extends across multiple physiological and developmental stages,
where they fine-tune gene expression networks with remarkable
precision [3]. By binding to complementary target mRNAs with high
specificity, miRNAs induce either transcript degradation through
cleavage or translational repression through inhibition of protein
synthesis, enabling precise spatial and temporal control over gene
expression. This sophisticated regulatory mechanism operates at
multiple levels, ensuring proper cellular function and organismal
development. This fine-tuning mechanism is particularly crucial for
plants as sessile organisms, allowing them to rapidly adjust their gene
expression profiles in response to internal cues and external stimuli.
The ability to modulate gene expression dynamically is essential for
plants to adapt to constantly changing environmental conditions
while simultaneously coordinating complex developmental
transitions that determine their growth patterns and reproductive
success [4-6].
The first miRNA, *lin-4*, was discovered in Caenorhabditis
elegans in 1993, revealing a novel layer of gene regulation mediated
by small RNAs. This groundbreaking finding challenged the
conventional understanding of genetic regulation and opened new
avenues in molecular biology. Subsequent research identified miRNAs
in animals and later in plants, demonstrating their evolutionary
conservation and functional significance. The discovery of the RNA
interference (RNAi) pathway further elucidated the mechanisms by
which small RNAs modulate gene expression, solidifying miRNAs as
key players in genetic regulation across eukaryotes [7-10].
The first plant miRNA, miR171, was identified in Arabidopsis
thaliana in 2002, marking a major milestone in plant molecular
biology. Early studies revealed that plant miRNAs differ from their
animal counterparts in their biogenesis, target specificity, and
functional roles. Unlike animal miRNAs, which often exhibit partial
complementarity to their targets, plant miRNAs typically bind with
near-perfect complementarity, leading to mRNA cleavage rather
than translational repression. The identification of conserved miRNA
families across land plants highlighted their fundamental roles in
development and stress responses. Advances in high-throughput
sequencing and bioinformatics have since expanded the catalogue
of known plant miRNAs, uncovering their extensive regulatory
networks [11-14].
This review provides a comprehensive overview of plant
miRNAs, beginning with their biogenesis and maturation processes.
We then discuss their critical functions in plant growth and
development, including their roles in shoot and root architecture,
leaf morphogenesis, and reproductive transitions. Additionally,
we examine how miRNAs mediate responses to abiotic and biotic
stresses, enabling plants to withstand adverse conditions. Finally, we
explore the evolutionary conservation of miRNAs across plant species
and their emerging applications in biotechnology, where engineered
miRNAs are being harnessed to enhance crop resilience and
productivity. By integrating current knowledge on miRNA biology,
this review underscores their significance in both fundamental plant
science and agricultural innovation.
miRNA biogenesis in plants
Transcription of Primary miRNAs (pri-miRNAs):
In plants, miRNA biogenesis begins with the transcription of
miRNA genes by RNA Polymerase II, producing long primary
transcripts called pri-miRNAs(Figure 1). These pri-miRNAs contain
stem-loop structures that are essential for subsequent processing.
Like typical mRNAs, they are modified with a 5’ cap and a 3’
polyadenylated tail, ensuring stability and proper nuclear processing.
The transcription of pri-miRNAs is tightly regulated, influenced
by developmental and environmental cues. This step ensures that
miRNA levels are finely tuned to meet cellular demands, laying the
foundation for downstream processing [15-18].Processing by DICER-LIKE (DCL) Proteins:
The pri-miRNAs are cleaved in the nucleus by the DICERLIKE1
(DCL1) protein complex, which generates precursor miRNAs
(pre-miRNAs) with shorter stem-loop structures. DCL1 works in
coordination with auxiliary proteins like HYL1 and SE to ensure
accurate and efficient processing. HYL1 stabilizes the pri-miRNADCL1
interaction, while SE aids in recruiting processing machinery
(Figure 1). The precise cleavage by DCL1 is crucial for producing
functional miRNA duplexes. Defects in these proteins can lead to
improper miRNA maturation, affecting plant growth and stress
responses [16-22].
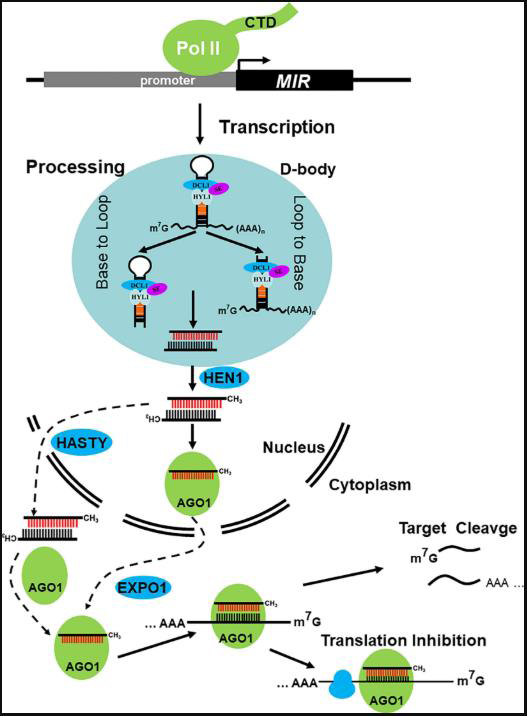
Figure 1:MIR genes are transcribed by RNA Polymerase II (Pol II) into
primary miRNA transcripts (pri-miRNAs). These pri-miRNAs are processed
into miRNA/miRNA* duplexes by the dicing complex in D-bodies, with
cleavage occurring in either the base-to-loop or loop-to-base direction. The
miRNA/miRNA* duplex is then methylated by the terminal methyltransferase
HEN1 in the nucleus. The mature miRNA is loaded into AGO1 and exported
via EXPO1, or the methylated duplex is transported to the cytoplasm via
HASTY before being incorporated into AGO proteins for RNA silencing.
Figure is adapted from Zhang et al. 2022 [16].
Nuclear Export and Maturation:
After processing, the miRNA duplex (miRNA:miRNA*) is
exported to the cytoplasm by HASTY, the plant homolog of exportin-5.
Once in the cytoplasm, the duplex is unwound, and the mature miRNA
(guide strand) is loaded into the RNA-induced silencing complex
(RISC). The passenger strand (miRNA*) is typically degraded, though
some may also play regulatory roles. The incorporation of the miRNA
into RISC marks the final step in maturation, enabling it to target
complementary mRNAs for silencing. This selective export ensures
only functional miRNAs mediate gene regulation [22-26].Mode of Action: mRNA Silencing:
Plant miRNAs primarily silence target mRNAs through cleavage,
mediated by Argonaute (AGO) proteins, particularly AGO1.
The miRNA-RISC complex binds near-perfect complementary
sequences, leading to mRNA degradation. Some miRNAs also repress
translation without cleavage, though this mechanism is less common
in plants. miRNA-mediated silencing regulates diverse processes,
including development, stress responses, and pathogen defense. The
precision of this system highlights its importance in maintaining
plant homeostasis and adaptability[27-31]Regulation of miRNA biogenesis
Transcription Factors (TFs):
Specific TFs, such as MYB, WRKY, and bZIP families, bind
to MIR gene promoters, either activating or repressing their
transcription in a tissue-specific or stress-dependent manner. For
example, in Arabidopsis, WRKY TFs modulate miRNA expression
during pathogen defense, while MYB factors regulate developmental
miRNAs. Some TFs act as master regulators, integrating hormonal and
environmental signals to control miRNA production. Additionally,
competing endogenous RNAs (ceRNAs) can sequester TFs, indirectly
influencing MIR gene expression [32-34].Epigenetic Modifications:
Chromatin structure profoundly impacts MIR gene expression,
with histone acetyltransferases (HATs) and methyltransferases
(HMTs) dynamically modifying nucleosome positioning. DNA
methylation at CpG islands, mediated by MET1 and DRM2, can
silence MIR loci, whereas demethylation activates them. In plants,
Polycomb Repressive Complex 2 (PRC2) deposits H3K27me3
marks to suppress certain MIR genes, while trithorax-group
proteins promote activation via H3K4me3. Environmental stresses,
such as cold or drought, can rapidly alter these epigenetic marks,
reprogramming miRNA expression [35-38].Drosha/DCL1 Complex: Post-Transcriptional Processing:
In animals, the microprocessor complex (Drosha-DGCR8)
recognizes and cleaves pri-miRNAs in the nucleus, whereas plants
use DCL1 in association with HYL1 and SERRATE for precise
processing. Structural features such as stem-loop stability and
flanking sequences determine cleavage efficiency. Mutations in these
core proteins lead to defective miRNA biogenesis, underscoring their
essential role. Auxiliary factors like TOUGH and DAWDLE further
enhance processing accuracy, ensuring proper miRNA maturation
[39-42].RNA Editing (ADAR/ADATs): Post-Transcriptional Processing:
Adenosine deaminases (ADARs) convert adenosine (A) to
inosine (I) in pri-miRNAs, altering their secondary structure and
potentially blocking Dicer cleavage. Similarly, cytidine deaminases
(e.g., APOBEC) induce C-to-U edits, which can disrupt miRNAmRNA
target pairing. These modifications are particularly prevalent
in neural and immune tissues, adding another layer of regulatory
complexity. In plants, RNA editing is less common but still influences
miRNA function under stress conditions [43-47].Alternative Splicing: Post-Transcriptional Processing:
Some MIR genes contain introns that undergo alternative
splicing, generating multiple pri-miRNA isoforms with distinct
hairpin structures. This can lead to the production of different
mature miRNAs from the same locus, expanding regulatory
diversity. For instance, splicing variants of MIR172 in Arabidopsis
produce functionally distinct miRNAs that regulate flowering time.
Dysregulation of splicing factors (e.g., SR proteins) can thus have
cascading effects on miRNA-mediated gene silencing [48-51].Abiotic Stress Responses:
Drought, extreme temperatures, and nutrient deficiencies trigger
kinase cascades (e.g., SnRK2, MAPKs) that phosphorylate miRNAprocessing
machinery, modulating their activity. For example,
osmotic stress induces SnRK2-mediated phosphorylation of DCL1,
enhancing miRNA production to suppress growth-related genes.
Heavy metals like cadmium can upregulate specific miRNAs (e.g.,
miR398) to activate detoxification pathways, illustrating adaptive
miRNA regulation [51-55].Biotic Stress Responses:
Pathogen infection activates immune signaling through PAMPtriggered
immunity (PTI), leading to miRNA reprogramming.
Salicylic acid (SA) and jasmonic acid (JA) pathways induce miRNAs
(e.g., miR393, miR160) that silence negative regulators of defense
responses. Viral suppressors of RNA silencing (VSRs) often target
DCL1 or AGO1 to block host miRNA biogenesis, highlighting
the evolutionary arms race between pathogens and host miRNA
machinery [56-61].RNA-Binding Proteins (RBPs):
Proteins like LIN28 bind to pre-miRNAs, inhibiting Drosha/
Dicer processing and promoting miRNA degradation. Conversely,
hnRNP A1 and KSRP stabilize pre-miRNAs, enhancing maturation.
In plants, DRB1 (HYL1) ensures accurate DCL1 cleavage, while
DRB2 fine-tunes miRNA abundance. RBPs also guide miRNAs to
specific subcellular locations, influencing their incorporation into
RISCs [62-66].Small RNA Stability Modifications:
3′-end methylation by HEN1 protects miRNAs from
exonucleolytic decay, a critical step in maintaining miRNA
longevity. Conversely, terminal uridylation (mediated by TUTases)
or adenylation marks miRNAs for degradation, providing a rapid
turnover mechanism. Environmental stresses can shift this balance;
for example, hypoxia increases uridylation of specific miRNAs,
reducing their stability and altering gene expression profiles [67-69].Functions in Plant Growth and Development
Leaf Development:
miRNAs play a crucial role in regulating leaf development by
controlling key transcription factors. For example, miR166 and
miR165 target HD-ZIP III family genes, which are essential for
establishing leaf polarity—determining the adaxial (upper) and abaxial
(lower) sides of leaves (Figure 2). Overexpression or suppression
of these miRNAs leads to abnormal leaf shapes, such as curled or
radialized leaves [70-72] Additionally, miRNAs like miR319 regulate
TCP TFs, influencing cell proliferation and leaf size. The precise
spatial and temporal expression of these miRNAs ensures proper
leaf morphogenesis during plant growth. Environmental factors
such as light and stress can modulate miRNA levels, further finetuning
leaf development [73-75]. The regulation of leaf development
by miR166/165 is crucial because HD-ZIP III transcription factors
control not just polarity but also vascular tissue formation. Without
proper miRNA-mediated control, leaves may develop improperly,
reducing photosynthesis efficiency. Additionally, these miRNAs help
plants adapt to environmental stresses by modulating leaf structure
under varying light conditions. Their role ensures balanced growth
between different leaf layers, optimizing light capture and gas
exchange[70-75]Root Architecture:
miRNAs are central to root development, particularly in lateral
root formation. The miR390-TAS3-ARF pathway is a key regulatory
module where miR390 triggers the production of trans-acting
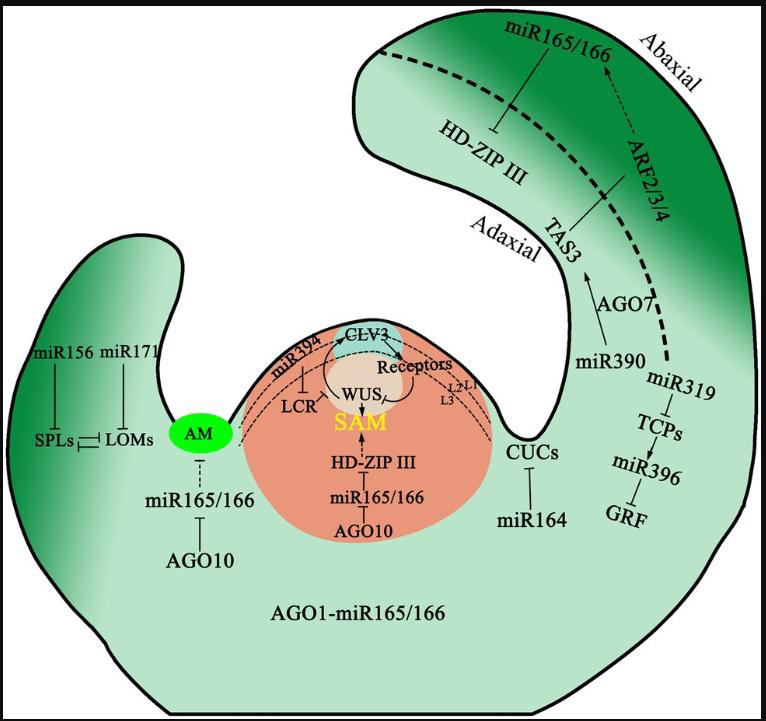
Figure 2:Regulatory network of small RNAs in shoot apical meristem
and leaf development. The diagram illustrates key small RNA-mediated
pathways controlling SAM maintenance, leaf polarity establishment, and
trichome initiation. miR394 represses LCR to activate WUS/CLV3 signaling,
while AGO10 sequesters miR165/166 to regulate meristem activity. HD-ZIP
III and ARF2/3/4 define adaxial-abaxial leaf domains under the control of
miR165/166 and TAS3 ta-siRNA, respectively. miR164, miR319/miR396,
and miR156/miR171 further modulate meristem initiation, leaf growth, and
trichome formation. Solid arrows indicate positive regulation; dashed lines
with perpendicular ends denote inhibitory interactions. Figure is adapted from
Dong et al. 2022 [72].
small interfering RNAs (tasiRNAs) from the TAS3 gene. These
tasiRNAs then suppress AUXIN RESPONSE FACTORs (ARFs),
particularly ARF2, ARF3, and ARF4, which are negative regulators
of lateral root growth (Figure 2). By modulating auxin signaling,
this pathway ensures proper root branching and soil exploration.
Mutations in this pathway result in altered root systems, affecting
nutrient uptake and plant stability. The miR390-TAS3-ARF pathway
is vital because auxin distribution dictates where lateral roots emerge,
improving nutrient and water absorption. Without this regulation,
roots may grow unevenly, weakening plant stability [72,76-79].
Juvenile-to-Adult Transition:
The transition from juvenile to adult vegetative phases is tightly
controlled by miR156, which targets SQUAMOSA PROMOTER
BINDING PROTEIN-LIKE (SPL) transcription factors. High levels
of miR156 in young plants suppress SPLs, maintaining juvenile traits
like leaf shape and delayed flowering. As the plant ages, miR156 levels
decline, releasing SPL repression and allowing the expression of adultphase
genes. Some SPLs also promote flowering, linking vegetative
phase change with reproductive timing. Environmental cues such as
light and temperature can influence miR156 accumulation, affecting
developmental timing. The gradual decline of miR156 ensures
that plants transition to maturity at the right time, preventing
premature flowering under unfavourable conditions. This regulation
is important because juvenile and adult leaves often have different
shapes and functions, affecting overall plant fitness. Environmental
factors like temperature can influence miR156 levels, allowing plants
to adjust their growth phases in response to seasonal changes [80-82].Floral Induction:
miR172 plays a pivotal role in promoting flowering by
repressing APETALA2 (AP2)-like transcription factors, which
act as floral repressors. As plants mature, miR172 levels increase,
reducing AP2-like activity and allowing floral meristem identity genes
(e.g., LFY, AP1) to be expressed. This regulatory switch ensures that
flowering occurs at the appropriate developmental stage. Some AP2-
like genes also regulate floral organ identity, making miR172 crucial
for both floral timing and patterning. By suppressing AP2-
like genes, miR172 ensures flowering occurs only when the plant has
sufficient energy and resources. This prevents wasted reproductive
efforts in poor growing conditions. Additionally, since AP2-like genes
also affect flower structure, miR172 indirectly ensures proper floral
organ development. Its role is critical for synchronizing flowering
with pollinators and optimal seed-setting conditions [83-86].Floral Organ Identity:
miRNAs contribute to floral patterning by regulating
key developmental genes. miR172, for instance, finetunes
AP2 expression, ensuring proper sepal and petal formation.
Another example is miR159, which targets MYB transcription factors
to control stamen development. Disruption of these miRNAs leads
to floral abnormalities, such as homeotic transformations (e.g., petals
turning into stamens). The precise spatiotemporal expression of these
miRNAs ensures correct floral organ specification. Proper floral
organ formation, controlled by miR172 and miR159, is essential for
successful pollination and seed production. If floral organs develop
incorrectly, pollination efficiency drops, reducing yield in crop plants.
These miRNAs also help maintain species-specific flower shapes,
which are often key for attracting the right pollinators [85-88].Seed Development:
During embryogenesis, miRNAs such as miR160 and miR167 regulate ARF genes to modulate
auxin signaling, which is critical for proper seed formation. miR160 targets ARF10/16/17, affecting embryo
patterning, while miR167 controls ARF6/8, influencing endosperm
development. Imbalances in these miRNAs can lead to seed
abortion or abnormal embryo morphology. Auxin-miRNA crosstalk
ensures coordinated seed growth and nutrient allocation.The role
of miR160 and miR167 in seed development is critical because
auxin signaling determines embryo orientation and nutrient flow.
Disruptions can lead to malformed seeds or even complete seed
abortion, affecting plant propagation. Since seeds are crucial for
the next generation, these miRNAs help maintain high germination
rates and seedling vigor, ensuring species survival [89-93]. (Table 1)
clearly indicates various functions of miRNAs in plant growth and
developmental processes.In the shoot apical meristem (SAM), miR394 is synthesized in the
protoderm and moves to subtending cells, where it represses LEAF
CURLING RESPONSIVENESS (LCR). This repression activates
WUSCHEL (WUS), maintaining stem cell identity and promoting
CLAVATA3 (CLV3) peptide expression. AGO10 specifically
sequesters miR165/166 in meristematic cells, counteracting its
activity to regulate SAM and axillary meristem (AM) development.
In contrast, AGO1 is broadly expressed in the apex and recruits
miR165/166 to form the RISC, ensuring proper meristem
function (Figure 2). During leaf primordia formation, HD-ZIP III
transcription factors are restricted to the adaxial (upper) side by
miR165/166, while ARF2/3/4 are confined to the abaxial (lower) side
via TAS3-derived trans-acting small interfering RNAs (ta-siRNAs).
Additionally, miR164 post-transcriptionally regulates two NAC domain
transcription factors, influencing embryonic meristem
initiation, boundary size control, and cotyledon establishment.
Leaf development is further modulated by miR319 and miR396,
which target TCP and GRF genes, respectively, coordinating cell
proliferation and differentiation. Meanwhile, miR156 and miR171
synergistically regulate trichome initiation by suppressing SPL and
LOM [70-93].
miRNAs in Stress Responses
Drought & Salinity (miR169-NF-YA Pathway):
miR169 plays a critical role in drought and salinity tolerance by
downregulating NF-YA transcription factors, which are involved in
stress-responsive gene expression. Under water-deficient conditions,
plants increase miR169 levels to suppress NF-YA, conserving energy
by reducing non-essential metabolic processes. This regulation helps
maintain cellular stability by preventing excessive stress-induced
damage. Additionally, miR169-mediated control ensures that only
essential stress-response genes are activated, improving survival rates
in harsh environments. Some crop plants genetically engineered to
overexpress miR169 show enhanced drought resistance, highlighting
its agronomic importance. The evolutionary conservation of miR169
across plant species suggests its fundamental role in abiotic stress
adaptation. Field studies indicate that natural variants with higher
miR169 expression perform better in arid regions, offering potential
for crop improvement programs[94-97].Nutrient Deficiency (miR399-PHO2 Regulation)
Under phosphate starvation, plants upregulate miR399, which
suppresses PHO2, a negative regulator of phosphate transporters. By
inhibiting PHO2, miR399 allows increased phosphate uptake from
the soil, ensuring proper growth even in low-nutrient conditions.
This miRNA-mediated regulation is crucial because phosphorus is
essential for ATP synthesis and nucleic acid formation. Interestingly,
miR399 is also transported from shoots to roots through the phloem,
coordinating systemic phosphate distribution. This mechanism
demonstrates how miRNAs help plants optimize nutrient use
efficiency under stress.Recent research shows that miR399 expression
patterns can serve as early indicators of phosphorus deficiency,
potentially enabling precision agriculture approaches. The discovery
of natural allelic variations in miR399 genes among crop wild relatives
may provide new genetic resources for breeding nutrient-efficient
varieties[98-101].
Pathogen Defense (miR393-Auxin Signaling):
When pathogens attack, plants elevate miR393 to suppress
auxin receptor genes (e.g., TIR1/AFB), reducing auxin signaling.
Since many pathogens exploit auxin pathways to weaken plant
immunity, miR393 acts as a defense mechanism by disrupting this
manipulation. The downregulation of auxin signaling also triggers
the activation of PR (Pathogenesis-Related) genes, enhancing
resistance. Studies show that plants with higher miR393 levels exhibit
stronger antibacterial and antifungal responses. This miRNA thus
serves as a molecular switch that prioritizes defense over growth
during infections.The speed of miR393 induction varies among plant
species, with faster responders showing greater disease resistance.
Agricultural applications could include developing miR393-
based biomarkers for early disease detection or engineering crops
with tunable miR393 expression for enhanced field resistance. In
Gossypium hirsutum, ghr-miR393-GhTIR1 module regulates plant
defense against Verticillium dahliaeby modulating auxin signaling.
Overexpression of ghr-miR393 or knockdown of GhTIR1 activates
ICS1 and NPR1, key components of SA-mediated defense. This
suppression of auxin signaling (via GhTIR1) enhances resistance by
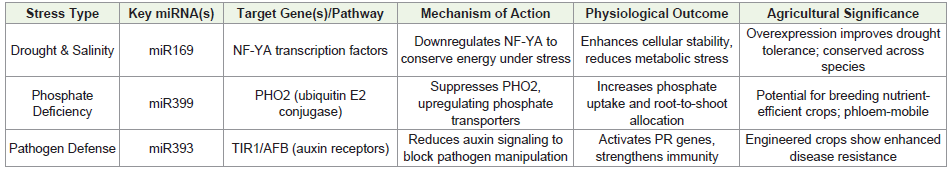
derepressing SA-dependent pathways (Figure 3)[102-104].(Table 2)
clearly indicates various roles of miRNAs in biotic and abiotic stress
responses.Evolutionary Conservation and Biotechnological Applications
Evolutionary Conservation of miRNAs:
The conservation of miRNAs like miR156 and miR172 across
land plants highlights their fundamental roles in regulating essential
biological processes. These miRNAs have been preserved over millions
of years of evolution, suggesting strong selective pressure to maintain
their functions in growth, development, and stress responses. Their
conserved sequences and target genes across diverse species indicate
a shared regulatory mechanism that has been fine-tuned through
evolutionary time. Studying these miRNAs provides insights into the
core genetic pathways that govern plant physiology. Additionally,
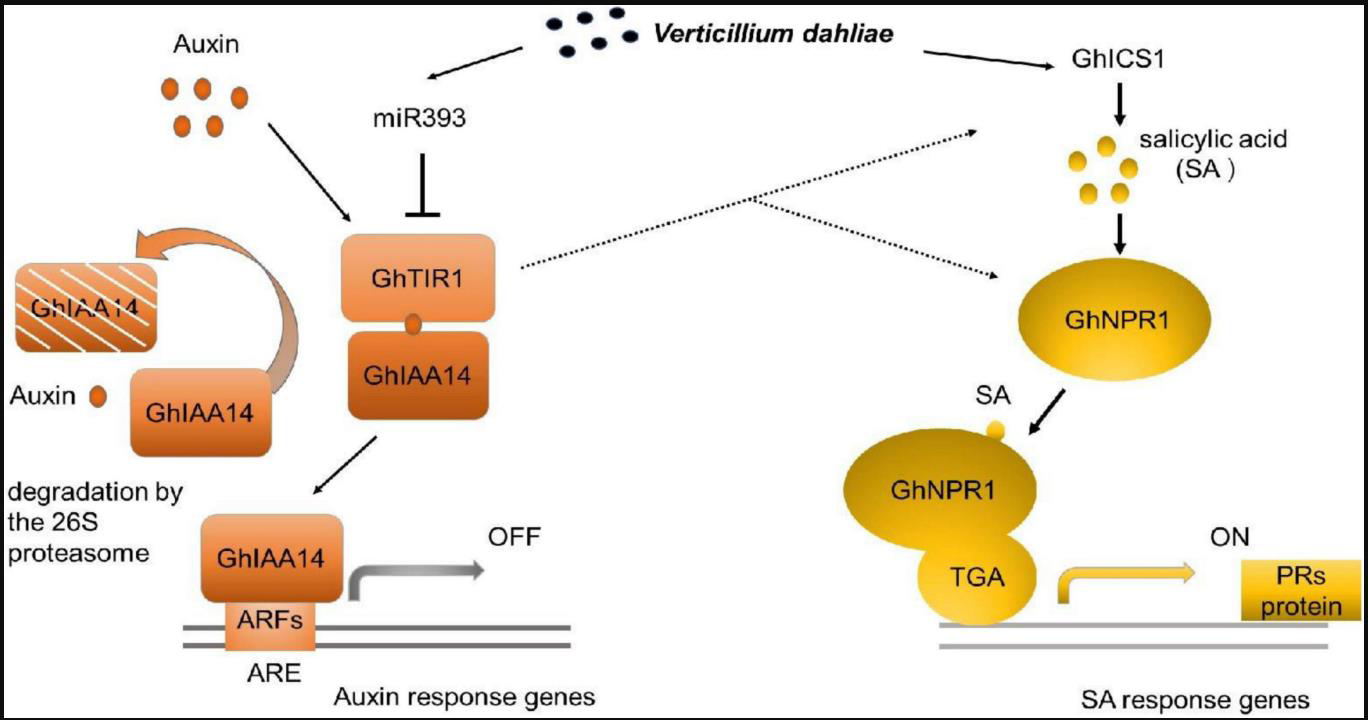
Figure 3:Model of ghr-miR393-GhTIR1 in plant immunity against Verticillium
dahliae. ghr-miR393 targets GhTIR1, suppressing auxin signaling and
activating ICS1-NPR1-mediated SA defense. Dashed lines indicate
enhanced ICS1/NPR1 expression upon ghr-miR393 overexpression or
GhTIR1 knockdown. ARE (auxin response element) may contribute to
pathway regulation. Figure is adapted from Shi et al. 2022 [104].
their conservation allows researchers to leverage knowledge
from model organisms to understand their roles in economically
important crops. The stability of these regulatory molecules across
species also suggests that manipulating them could have predictable
and widespread effects in plant biotechnology[105-107].
Biotechnological Applications of Engineered miRNAs:
Engineered miRNAs offer a powerful tool for precise genetic
modification in crops, enabling targeted enhancement of stress
tolerance and yield. By designing artificial miRNAs, scientists can
silence or modulate specific genes involved in stress responses,
nutrient utilization, or developmental pathways without introducing
foreign proteins. This approach reduces unintended effects compared
to traditional transgenic methods. Engineered miRNAs can be
tailored to fine-tune gene expression, optimizing traits such as
drought resistance, disease immunity, or flowering time. Their
small size and high specificity make them easier to incorporate into
plant genomes while minimizing regulatory concerns. Furthermore,
since endogenous miRNA pathways are already present in plants,
engineered miRNAs integrate seamlessly into existing regulatory
networks. This technology holds great promise for sustainable
agriculture by improving crop resilience and productivity under
challenging environmental conditions [108-110].Future Perspectives
Expanding miRNA Discovery Through Single-Cell Sequencing:
Future research should leverage single-cell RNA sequencing to
uncover cell-type-specific miRNA expression patterns, providingunprecedented resolution in understanding developmental and stress
responses. This approach will reveal how miRNAs fine-tune gene
regulation in individual cell types, such as root hairs or guard cells,
under varying conditions. Integrating spatial transcriptomics could
further map miRNA activity across tissues, enhancing our knowledge
of their localized functions. Such advancements will enable the design
of precision-engineered miRNAs for targeted crop improvement.
Deciphering miRNA Crosstalk with Epigenetic Mechanisms:
Exploring the interplay between miRNAs and epigenetic
modifications (DNA methylation, histone marks) will uncover new
regulatory layers in stress adaptation. Future studies should investigate
how environmental cues alter miRNA expression via chromatin
remodeling and how these changes are inherited. Understanding this
crosstalk could lead to epigenetic editing strategies that enhance stress
memory in crops. Additionally, identifying miRNAs that regulate
epigenetic modifiers may reveal novel targets for biotechnology
applications.Developing miRNA-Based Biomarkers for Precision Agriculture:
miRNA expression profiles could serve as early diagnostic
biomarkers for stress conditions, nutrient deficiencies, or disease
susceptibility. Future work should focus on field-deployable detection
methods, such as portable PCR or nanosensors, to monitor miRNA
dynamics in real time. This could enable preemptive agricultural
interventions, optimize resource use and minimize yield losses.
Machine learning models trained on miRNA expression data may
further improve predictive accuracy for crop management.Engineering Synthetic miRNA Networks for Climate Resilience:
Advancements in synthetic biology could allow the design of
artificial miRNA circuits that dynamically respond to environmental
triggers (e.g., drought, heat). Future efforts should focus on creating
feedback-regulated miRNA systems that fine-tune stress responses
without compromising growth. Combining multiple engineered
miRNAs into synergistic networks may enhance multi-stress
tolerance. Field trials of such designs will be critical to assess their
efficacy under real-world conditions.Harnessing miRNA-Mediated RNAi for Pest and Pathogen Control:
Future applications could exploit miRNA pathways to develop
RNAi-based biopesticides that target herbivores or pathogens while
sparing beneficial organisms. Research should optimize delivery
methods, such as nanoparticle carriers or root uptake, to ensure
stability and specificity. Engineered miRNAs could also silence
virulence genes in pathogens, offering a sustainable alternative to
chemical pesticides. Regulatory frameworks must evolve to address
the ecological implications of such technologies.Exploring Horizontal miRNA Transfer in Plant-Microbe Interactions:
Emerging evidence suggests miRNAs may be exchanged between
plants and associated microbes, influencing symbiosis or defense.
Future studies should investigate the mechanisms and functional
consequences of this cross-kingdom communication. Understanding
how microbial miRNAs modulate host gene expression could lead to
novel biofertilizers or biocontrol agents. This field may uncover new
dimensions of plant-microbe coevolution.Integrating miRNA Editing with CRISPR-Cas Technologies:
Combining CRISPR-based genome editing with miRNA
manipulation could enable simultaneous tuning of multiple gene
networks. Future research should develop tools for precise miRNA
gene editing (e.g., promoter modifications, stem-loop alterations) to
optimize expression levels. Dual-function systems, where CRISPR
guides and miRNAs target complementary pathways, may enhance
trait stacking in crops. Ethical and regulatory considerations will be
paramount in deploying such advanced technologies.Bridging miRNA Research with Crop Wild Relatives for Breeding:
Future breeding programs should mine miRNA diversity in
crop wild relatives to identify natural alleles associated with stress
resilience. Comparative genomics and pan-miRNAome analyses
could reveal conserved and species-specific regulatory nodes.
Introgression of beneficial miRNA variants via marker-assisted
selection may accelerate the development of climate-smart crops. This
approach aligns with sustainable agriculture by reducing reliance on
transgenic modifications.Conclusion
MicroRNAs (miRNAs) play vital roles in plant biology,
regulating gene expression through mRNA cleavage or translational
repression to control growth, development, and stress responses.
Their evolutionary conservation highlights their importance across
species, while biotechnological advances demonstrate their potential
for engineering stress-resistant crops. This review explores miRNAmediated
regulation, emphasizing their role in plant physiology and
agricultural innovation. Emerging technologies, such as single-cell
sequencing and synthetic miRNA networks, may further enhance
crop resilience and productivity. Understanding miRNAs is key to
addressing global food security through precision breeding and
biotechnology.
References
Citation
Netala VR, Al Aboud NM, Tartte V. MicroRNAs as Master Regulators of Plant Development and Stress Adaptation. J Plant Sci Res. 2025;12(2): 281